Project Lead: Siva Kasinathan, UW School of Medicine
eScience Liaisons: Andrew Fiore-Gartland, Bryna Hazelton
Despite an explosion in DNA sequencing technology, many genome projects, including the Human Genome Project, remain fundamentally unfinished. Gaps in genome assemblies occur in regions composed of repeated sequences. Human centromeres, which are loci that ensure proper partitioning of genetic material at each cell division, are one such class of unassembled sequence and account for an estimated 60 million base pairs of a genome that is 3 billion base pairs in length. Centromere dysfunction may be associated with cancers and developmental disorders such as Down syndrome; however, the inability to exactingly interrogate centromere sequence has impeded a clear understanding of centromere biology in human health and disease.
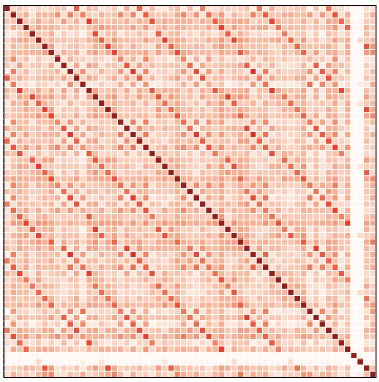
Gene sequencing is carried out by ‘reading’ chunks of the genome at a time, and then piecing those chunks back together, much like putting together the pieces of a jigsaw puzzle. Unfortunately, regions of the genome that contain a large number of repeated patterns are particularly difficult to reassemble. This incubator project is focused on developing methods for trying to reassemble these parts of the genome. In the first half of this incubator period we developed a ‘fake’ genome which would allow us to test which methods have the potential to be successful and examined whether piecing together sequences based on cross-correlations patterns is likely to be effective.